Understanding the mechanisms underlying infectious diseases is fundamental to develop prevention strategies. Host-Pathogen Interactions (HPI), which includes from the initial invasion of host cells by the pathogen through the proliferation of pathogen in their host, are studied to find potential genomic targets for the development of novel drugs, vaccines, and other therapeutics. Determining which proteins are involved in the interaction system behind an infectious process is the first step to develop an efficient disease control strategy.
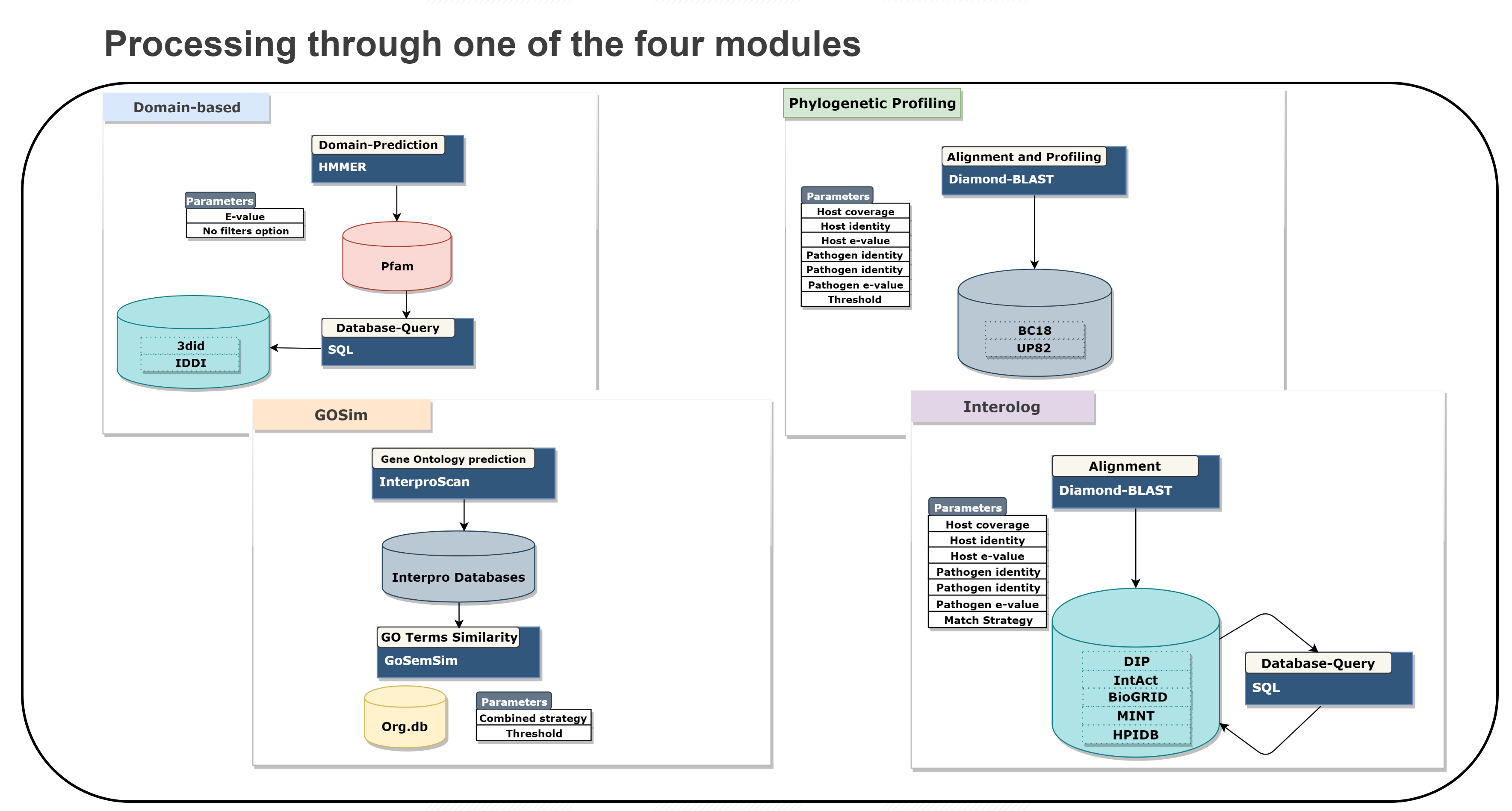
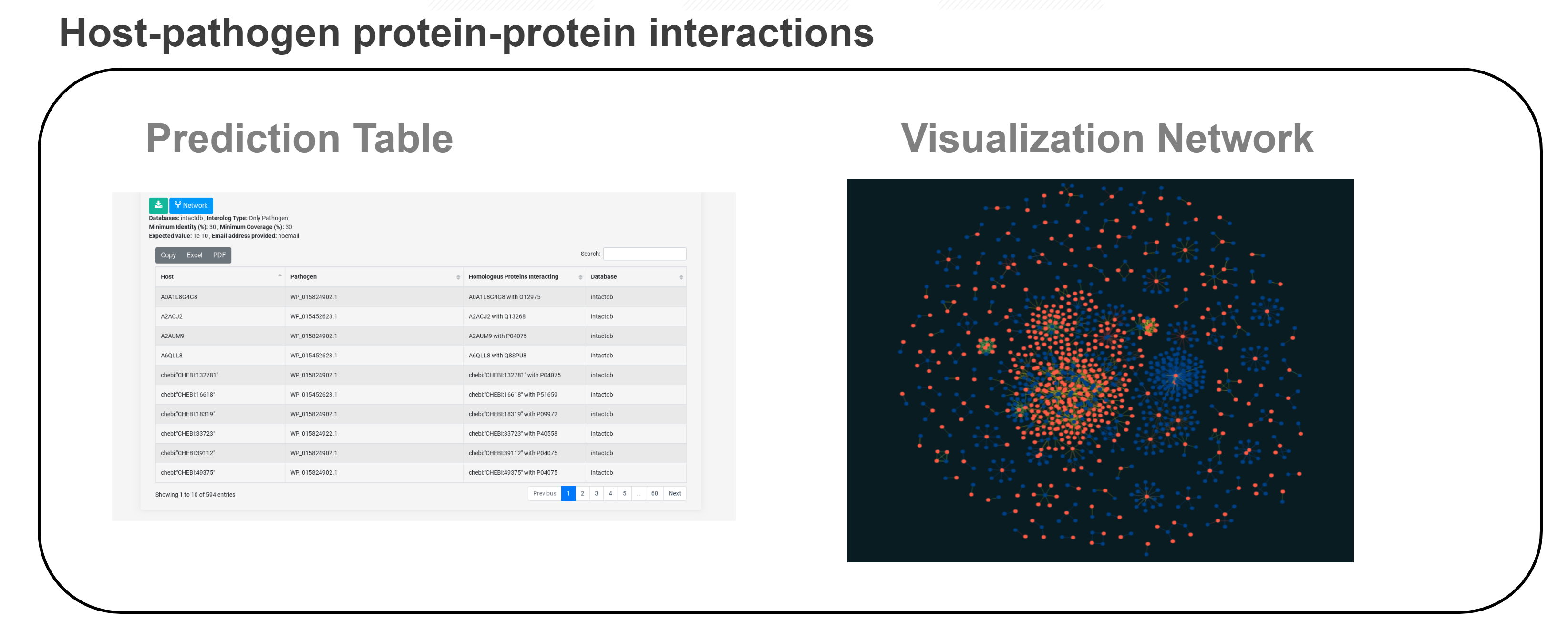
In silico prediction methods have barely been implemented as web services to infer novel HPIs, and there is not a single framework which combines several of those approaches to produce and visualize a comprehensive analysis of host-pathogen interactions. We introduce PredHPI, a powerful framework that integrates both the detection and visualization of interaction networks in a single web service, facilitating the apprehension of model and non-model host-pathogen systems to aid the biologists in building hypotheses and designing appropriate experiments. PredHPI is built on high-performance computing resources on the backend capable of handling proteome-scale data from both host as well as pathogen. Data are displayed in an information-rich and interactive visualization, which can be further customized with user-defined layouts.